By aggregating self-reported health statuses across millions of users, we seek to characterize the variety of health information discussed in Twitter. We describe a topic modeling framework for discovering health topics in Twitter, a social media website. This is an exploratory approach with the goal of understanding what health topics are commonly discussed in social media. This paper describes in detail a statistical topic model created for this purpose, the Ailment Topic Aspect Model (ATAM), as well as our system for filtering general Twitter data based on health keywords and supervised classification. We show how ATAM and other topic models can automatically infer health topics in 144 million Twitter messages from 2011 to 2013. ATAM discovered 13 coherent clusters of Twitter messages, some of which correlate with seasonal influenza (r = 0.689) and allergies (r = 0.810) temporal surveillance data, as well as exercise (r = .534) and obesity (r = −.631) related geographic survey data in the United States. These results demonstrate that it is possible to automatically discover topics that attain statistically significant correlations with ground truth data, despite using minimal human supervision and no historical data to train the model, in contrast to prior work. Additionally, these results demonstrate that a single general-purpose model can identify many different health topics in social media.
FiguresCitation:Paul MJ, Dredze M (2014) Discovering Health Topics in Social Media Using Topic Models. PLoS ONE 9(8): e103408. doi:10.1371/journal.pone.0103408
Editor: Renaud Lambiotte, University of Namur, Belgium
Received: January 7, 2014; Accepted: July 2, 2014; Published: August 1, 2014
Copyright: © 2014 Paul, Dredze. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Funding:Mr. Paul was supported in part by a National Science Foundation Graduate Research Fellowship under Grant No. DGE-0707427 and a PhD fellowship from Microsoft Research. Publication of this article was funded in part by the Open Access Promotion Fund of the Johns Hopkins University Libraries. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Competing interests: Dr. Dredze reports receipt of compensation for travel for talks at various academic, corporate, and governmental entities and consulting for Directing Medicine, Progeny Systems, and Sickweather. Mr. Paul serves on the advisory board for Sickweather. This does not alter the authors' adherence to PLOS ONE policies on sharing data and materials.
IntroductionSeveral studies have utilized social media for tracking trends and analyzing real world events, including news events, [1] natural disasters, [2] user sentiment, [3] and political opinions. [4]–[5]Twitter is an especially compelling source of social media data, with over half a billion user-generated status messages (“tweets”) posted every day, often publicly and easily accessible with streaming tools. [6] By aggregating the words used by millions of people to express what they are doing and thinking, automated systems can approximately infer what is happening around the world. Researchers have begun to tap into social media feeds to monitor and study health issues, [7] with applications in disease surveillance and other epidemiological analysis.
By far the most commonly analyzed disease in social media is influenza. Many researchers have tracked influenza in social media data, most commonly Twitter, using a variety of techniques such as linear regression, [8]–[10] supervised classification, [11]–[12] and social network analysis. [13] Researchers have also used social media to study cholera, [14] dental pain, [15] and cardiac arrest, [16] as well as population behavior including physical activities,[17] mood and mental health, [18]–[19] and alcohol, [9], [20] tobacco, [21] and drug use. [22]Twitter has a desirable property of being a real time data source, in contrast to surveys and surveillance networks that can take weeks or even years to deliver information. Additionally, users of Twitter may candidly share information that they do not provide to their doctor, and thus it is potentially a source of new information, such as off-label use of medications. [23], [24].
Studies like these rely on the detection of specific illnesses such as influenza or health topics such as exercise. In this work, we instead describe how to perform discovery of ailments and health topics. We do this using topic models, which automatically infer interesting patterns in large text corpora. We believe an exploratory, discovery-driven approach can serve us a useful starting point for medical data mining of social media, by automatically identifying and characterizing the health topics that are prominently discussed on social media. Our goal is not to improve modeling of any one specific illness, but to demonstrate a model for illness discovery. While we may validate the discovered illnesses against specialized approaches for tracking each specific illness, the strength of our model is that it allows discovery of new illness in new data without a priori knowledge. Furthermore, our list of discovered illnesses contains several that have previously been unexplored in Twitter, suggesting new areas for directed research, described in the Discussion section.
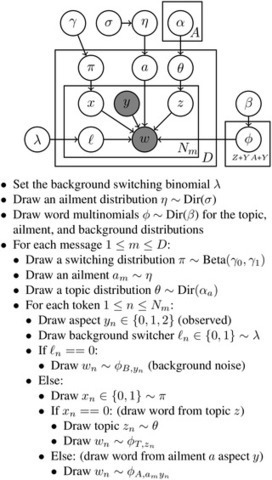
In this paper, we describe a statistical topic modeling framework for identifying general public health information from millions of health-related tweets. In addition to a basic topic model, we also describe our Ailment Topic Aspect Model (ATAM), previously used to analyze tweets from 2009–10. [24] This framework is used to explore the diversity of health topics that are discussed on Twitter, and we find that many health topics correlate with existing survey data. Our specific contributions are: (1) we describe a current end-to-end framework for data collection and analysis, which includes multiple data streams, keyword filters, and supervised classifiers for identifying relevant data; (2) we analyze a set of 144 million health-related tweets that we have been downloading continuously since August 2011; (3) we provide many previously unpublished details about the creation of our classifier for identifying health tweets and details of ATAM, our specialized health topic model, including procedures for large-scale inference; (4) we evaluate this framework and topic model quality by comparing temporal and geographic trends in the data with external data sources. We experiment with both a basic topic model and ATAM, as well as individual keyword filters for comparison. This article is an extension of an earlier unpublished technical report [25] and includes a longer explanation of ATAM and LDA, more technical detail such as the Gibbs sampling update equations, and more experimental comparisons between various approaches than any of our previous studies on this subject.
Via Plus91





 Your new post is loading...
Your new post is loading...